ProMetheus
For unravelling the molecular response to chemicals and for detecting mechanisms underlying deleterious phenotypes of cells, organs and organisms, proteomics and metabolomics are well established tools, since they are characterising actual molecular functions.
We established the ProMetheus platform for mass spectrometry-based (meta)proteomics and metabolomics in 2008. Prometheus, the titan in Greek mythology who brought the fire to mankind, was the inspiration for our mission. We develop and provide state-of-the-art tools to shed light on research questions for which no or not sufficient insights are available so far. In addition to constant method improvement and development, our research focuses on the effects of chemicals and, in particular, on microbiome-host interactions in health and disease. In ProMetheus, we have established untargeted and targeted measurement of proteins and metabolites, metaproteomics, and novel data evaluation workflows for in-house and external use. Another strength of the platform is our expertise in metaproteomics, which can strongly support the functional analysis of effects. Our research is closely connected to other groups within the UFZ and beyond, and we are open to further collaborations.
Targeted metabolomics approach for characterising cellular, systemic and microbial metabolism
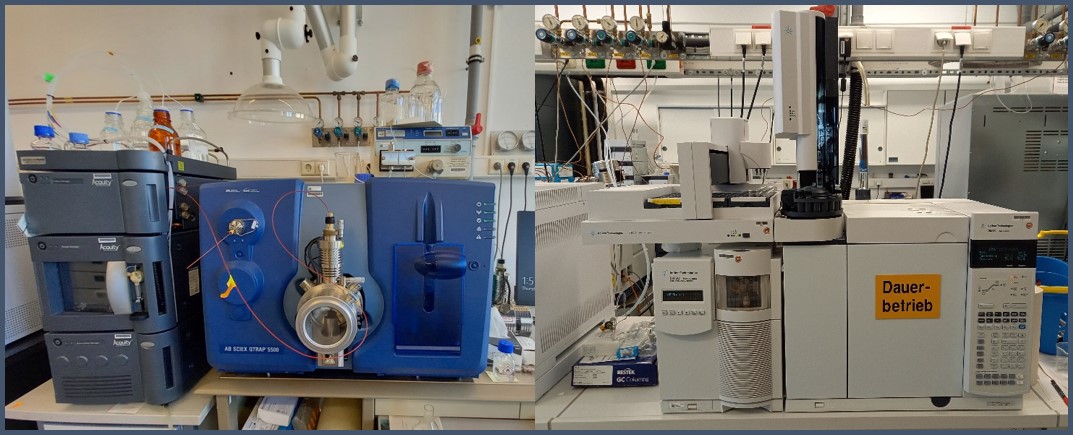
People are exposed to a wide range of chemicals in the environment. Beside other effects these can lead to changes in metabolism, which in turn can lead to functional disorders and even to the development of disease. The analyses cover a wide range of topics, from the investigation of the host-microbiome interaction to effects of chemicals on organ or cellular level. Targeted metabolomics is able to accurately identify both the chemical compounds as well as the endogenous metabolites. And, when calibration standards are used, mass spectrometry is well suited to absolutely quantify the targeted analytes. For our analyses we use mass spectrometry, coupled to liquid or gas chromatography. Due to the many classes of substances that can be responsible for functional disorders, we adapt our methods and are steadily developing new ones. The state of the art equipment allows us to select the best solution in respect to the type of mass spectrometer. This includes 3 LC-MS and 2 GC-MS systems from Sciex, 2 Agilent QToF systems and an Orbitrap IDX which is currently being procured.

Untargeted metabolomics for characterizing microbiome host interactions
The focus of the group is to reveal metabolic crosstalk between the host and microbiome, as well as the metabolic exchange within the microbiome. To detect and identify metabolites in an unbiased manner or to identify unknown compounds untargeted metabolomic approaches are a valuable tool. In this method we are able to screen the entire metabolome of a specific sample type such as body fluids, tissue samples, bacterial and cellular model system and environmental samples. The advantage of this method lies in the possibility to get a global and comprehensive analysis of the sample without predetermining which metabolites should be measured. The resulting data can be used as a metabolic fingerprint and also be used as a basis for deciding which targeted metabolomic approaches should be developed. For this approach we use HPLC-HR-MS/MS (2 Agilent ToF systems and an Orbitrap IDX).
Isotopic flux for identifying effects of chemicals on metabolic regulation
Metabolic pathways and transformation kinetics can be changed or disturbed by the exposure to toxins and ubiquitous chemicals. Isotopic tracer analysis allows an isotopic labelled metabolic substrate to be followed through downstream metabolism. By following the tracers we are able to determine which metabolic pathways are utilized. In combination with time series measurements this allows us to get insights into the kinetics of the metabolic transformations. Commonly used for tracing are labelled carbon (13C), nitrogen (15C) or hydrogen (2H) sources, and also combination of multiple tracers can be introduced. For that purpose we use targeted LC-MS/MS (QTRAP Sciex System) and we plan to extend the isotopic tracer measurements to the newly purchased Orbitrap IDX System.
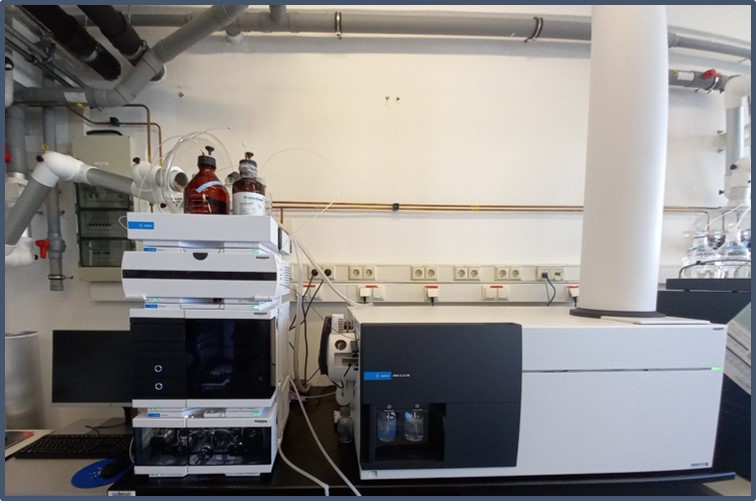

Cellular and systemic proteomics for characterising effects of environmental contaminants
The public is increasingly exposed to a variety of environmental contaminants, which have the potential to cause adverse effects. To date, risk assessment relies mainly on in vivo experiments, which are costly and ethically questionable. Our aim is to investigate environmental contaminants in vitro and thus contribute to the 3Rs principle (replace, reduce, refine) of animal testing. Thereby, we focus on the effects on the immune system as a barrier against foreign substances. The immune system can regulate the microbiome homeostasis and can cause severe diseases in the case of dysbiosis, such as chronic inflammation that cannot be resolved. In addition to studying toxicological endpoints, we use targeted and untargeted proteomics to gain insights into the modes of action of environmental contaminants. Besides information on protein abundances, we can also use untargeted proteomics to investigate the effects on post-translational modifications, which provide further detailed insights into the regulation of pathways.
The instruments used for these studies, a QExactive HF and a QTRAP 5500, are part of the ProMetheus platform. The ProMetheus platform thus provides the basis for analysing modes of action of environmental contaminants, facilitating the definition of adverse outcome pathways (AOPs) and thus future risk assessment in accordance with the 3Rs principle of animal testing.
Microbial (Meta)Proteomics for unravelling microbiome-host interaction
Microbial communities (microbiomes) are major drivers in nutrient cycles and key factors in human health. High-throughput genome sequencing technologies have revealed the immense variety of microbes and the complexity of microbiomes. They are essential for the functioning of natural health. Many cellular functions (metabolism, transport, stress response) are represented by proteins. Improving human health, especially to unravel microbiome-host interactions, requires a detailed understanding of microbes at the functional level. Metaproteomics, detecting expressed proteins, covers the functional entity of microbiomes and complements metagenomics and metatranscriptomics.
We are using mass spectrometry-based metaproteomics (Q Exactive HF instrument) in the ProMetheus platform to analyse the metaproteomes describing microbiome-host interaction thus gaining a deeper understanding of this interaction. The method is increasingly becoming the focus of research and will become an indispensable tool within the next 10 years.
