Analytical methods
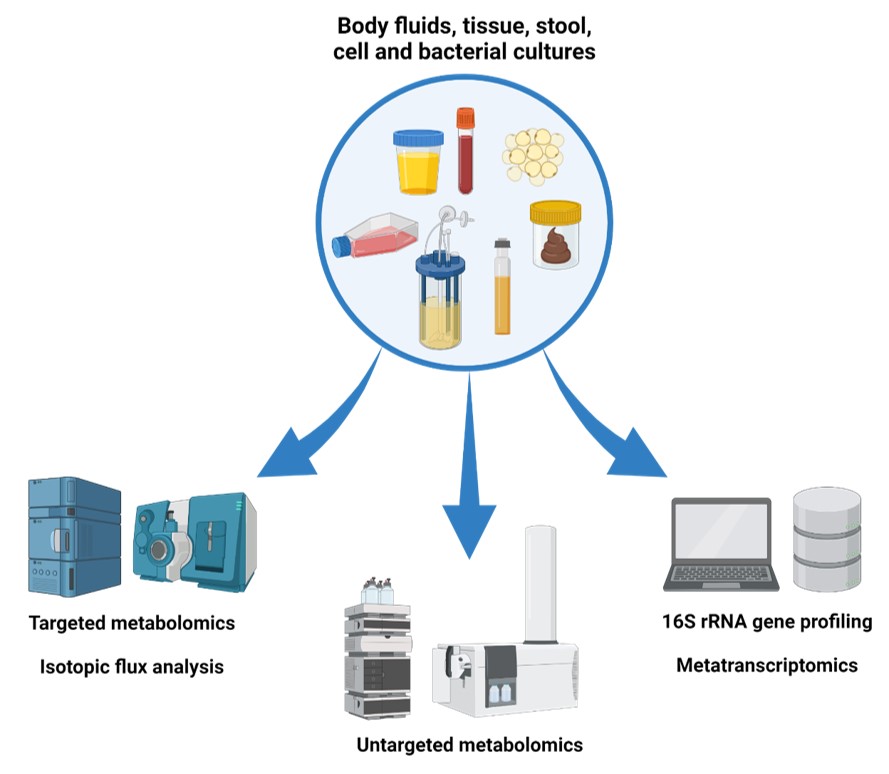
- Targeted metabolomics can accurately identify specific known metabolites and quantify them. In our working group we use high pressure liquid chromatography coupled to multi-reaction-monitoring mass spectroscopy for analysis (more detail here).
- Isotopic tracer analysis allows a isotopic labelled metabolic substrate to be followed through downstream metabolism. Commonly used for tracing are labelled carbon (13C), nitrogen (15N) or hydrogen (2H) sources (more detail here).
- Untargeted metabolomics allows a global screening of known and unknown metabolites in one sample. In our working group we have access to a high-performance liquid chromatography coupled to a time-of-flight or orbitrap mass spectrometer (more detail here).
- 16S rRNA gene profiling is a common method used to determine relative abundances of bacterial taxa in microbial communities. It is based on the sequencing and relative quantification of the 16S rRNA gene ubiquitous to all bacteria (more detail here).
- Metatranscriptomics is a powerful method to determine taxonomic structure and functions expressed by microbial communities. The method is based on the analysis of the entire set of transcripts from the community (more detail here).
