Research Focus:
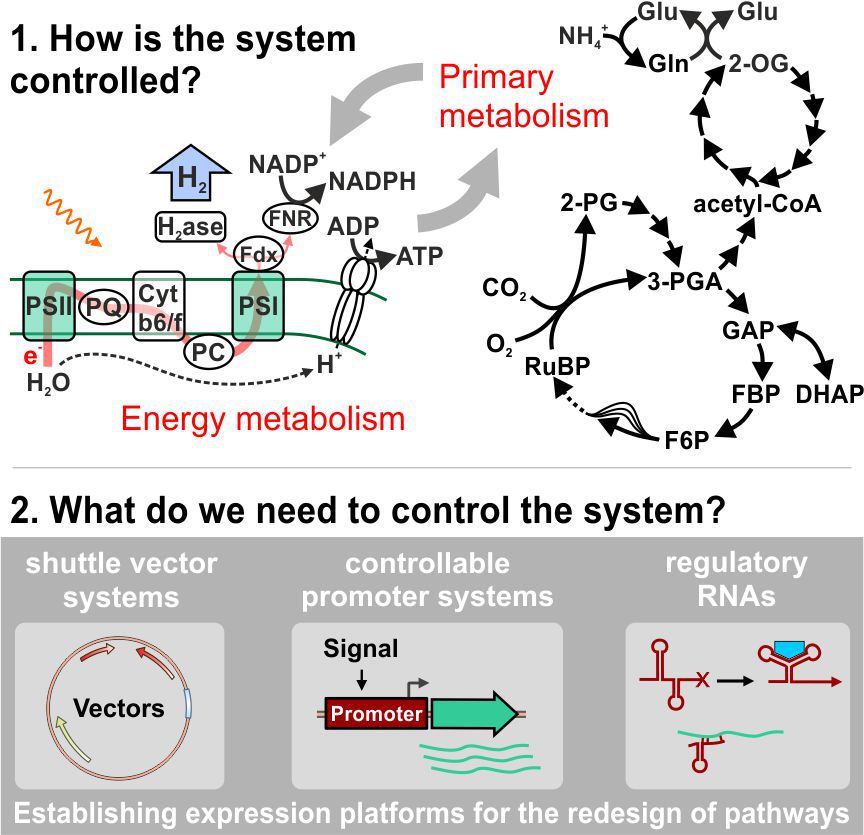
Our research is settled at the interspace of basic and applied research in the fields of genetics and microbiology with a major focus on the phylum of cyanobacteria. Cyanobacteria are the only prokaryotes performing oxygenic photosynthesis, a process which uses light energy and CO2 to form carbohydrates and biomass. Due to their photosynthetic lifestyle there is a rising interest for cyanobacteria in green biotechnology. Accordingly, an in-depth understanding of their photosynthetic apparatus as well as regulatory mechanisms that control electron fluxes to the coupled primary metabolism is of fundamental interest. Hence, our research aims at the basic understanding of those processes that control photosynthesis and metabolism in this interesting bacterial group. The obtained knowledge will provide a scientific fundament for the design of photosynthesis-driven biotechnological applications, e.g. the sustainable production of hydrogen which is a natural byproduct of photosynthesis in several cyanobacteria.
In addition, albeit the biotechnological potential of cyanobacteria is more and more exploited, one of the main problems still compromising their application is the rather limited availability of molecular toolboxes for genetic engineering. This includes the lack of easily controllable promoters that allow a wide range of expression levels for heterologous genes (e.g. for designed pathways that produce valuable chemicals). In addition to these expression platforms there is a high demand for tools that allow the integration of the engineered genetic information into the desired cyanobacterial host strains. Therefore, we also aim to increase the number of molecular toolboxes, i.e. regulatory systems for the utilization in cyanobacteria.
Group Leader:
PhD-Students:
Technicians:
Students:
Lara Kanzog
Carla Klemm
Index:
You could use our publication index for further requests.
2025 (3)
- Itzenhäuser, M.A., Enkerlin, A.M., Dewald, J.A., Avşar, B., Stauder, R., Halpick, H., Schaale, R., Baumann, L.M., Fernandez Merayo, N., Maskow, T., Selim, K.A., Weinberg, C.E., Klähn, S. (2025):
Deciphering guanidine assimilation and riboswitch-based gene regulation in cyanobacteria for synthetic biology applications
Proc. Natl. Acad. Sci. U.S.A. 122 (49), e2519335122
10.1073/pnas.2519335122 - Lupacchini, S., Stauder, R., Opel, F., Klähn, S., Schmid, A., Bühler, B., Toepel, J. (2025):
Co-expression of auxiliary genes enhances the activity of a heterologous O2-tolerant hydrogenase in the cyanobacterium Synechocystis sp. PCC 6803
Biotechnol. Biofuels Bioprod. 18 , art. 41
10.1186/s13068-025-02634-5 - Veit, M.C., Stauder, R., Bai, Y., Gabhrani, R., Schmidt, M., Klähn, S., Lai, B. (2025):
The necessity of multi-parameter normalization in cyanobacterial research: A case study of the PsbU in Synechocystis sp. PCC 6803 using CRISPRi
J. Biol. Chem. 301 (11), art. 110763
10.1016/j.jbc.2025.110763
2024 (2)
- Klähn, S. (2024):
Natürlicher Inhibitor der Nitritreduktase steuert Nitritsekretion in Cyanobakterien [Natural inhibitor of nitrite reductase controls nitrite secretion in cyanobacteria]
Biospektrum 30 (4), 423 - 423
10.1007/s12268-024-2234-6 - Klähn, S., Opel, F., Hess, W.R. (2024):
Customized molecular tools to strengthen metabolic engineering of cyanobacteria
Green Carbon 2 (2), 149 - 163
10.1016/j.greenca.2024.05.002
2023 (4)
- Biedermann, P., Engelmann, S., Klähn, S., Seiffert-Störiko, A., Sieben, C., Sander, J., Kruck, D. (2023):
Journal Club
Biospektrum 29 (4), 382 - 385
10.1007/s12268-023-1963-2 - Opel, F., Axmann, I.M., Klähn, S. (2023):
The molecular toolset and techniques required to build cyanobacterial cell factories
In: Bühler, K., Lindberg, P. (eds.)
Cyanobacteria in biotechnology. Applications and quantitative perspectives
Adv. Biochem. Eng. Biotechnol. 183
Springer Nature, p. 65 - 103
10.1007/10_2022_210 - Opel, F., Itzenhäuser, M.A., Wehner, I., Lupacchini, S., Lauterbach, L., Lenz, O., Klähn, S. (2023):
Towards a synthetic hydrogen sensor in cyanobacteria: Functional production of an oxygen-tolerant regulatory hydrogenase in Synechocystis sp. PCC 6803
Front. Microbiol. 14 , art. 1122078
10.3389/fmicb.2023.1122078 - Toepel, J., Karande, R., Klähn, S., Bühler, B. (2023):
Cyanobacteria as whole-cell factories: current status and future prospectives
Curr. Opin. Biotechnol. 80 , art. 102892
10.1016/j.copbio.2023.102892
2022 (5)
- Bolay, P., Hemm, L., Florencio, F.J., Hess, W.R., Muro-Pastor, M.I., Klähn, S. (2022):
The sRNA NsiR4 fine-tunes arginine synthesis in the cyanobacterium Synechocystis sp. PCC 6803 by post-transcriptional regulation of PirA
RNA Biol. 19 (1), 811 - 818
10.1080/15476286.2022.2082147 - Bolay, P., Schlüter, S., Grimm, S., Riediger, M., Hess, W.R., Klähn, S. (2022):
The transcriptional regulator RbcR controls ribulose-1,5-bisphosphate carboxylase/oxygenase (RuBisCO) genes in the cyanobacterium Synechocystis sp. PCC 6803
New Phytol. 235 (2), 432 - 445
10.1111/nph.18139 - Bozan, M., Popp, D., Kallies, R., Nunes da Rocha, U., Klähn, S., Bühler, K. (2022):
Whole-genome sequence of the filamentous diazotrophic cyanobacterium Tolypothrix sp. PCC 7712 and its comparison with non-diazotrophic Tolypothrix sp. PCC 7601
Front. Microbiol. 13 , art. 1042437
10.3389/fmicb.2022.1042437 - Brandenburg, F., Klähn, S., Schmid, A., Krömer, J.O. (2022):
Produktion von Aminosäurederivaten in Cyanobakterien [Production of amino acid derivatives in cyanobacteria]
Biospektrum 28 (3), 341 - 343
10.1007/s12268-022-1756-z - Opel, F., Siebert, N.A., Klatt, S., Tüllinghoff, A., Hantke, J.G., Toepel, J., Bühler, B., Nürnberg, D.J., Klähn, S. (2022):
Generation of synthetic shuttle vectors enabling modular genetic engineering of cyanobacteria
ACS Synth. Biol. 11 (5), 1758 - 1771
10.1021/acssynbio.1c00605
2021 (6)
- Bolay, P., Muro-Pastor, M.I., Rozbeh, R., Timm, S., Hagemann, M., Florencio, F.J., Forchhammer, K., Klähn, S. (2021):
The novel PII-interacting protein PirA controls flux into the cyanobacterial ornithine-ammonia cycle
mBio 12 (2), e00229-21
10.1128/mBio.00229-21 - Brandenburg, F., Theodosiou, E., Bertelmann, C., Grund, M., Klähn, S., Schmid, A., Krömer, J.O. (2021):
Trans-4-hydroxy-L-proline production by the cyanobacterium Synechocystis sp. PCC 6803
Metab. Eng. Commun. 12 , e00155
10.1016/j.mec.2020.e00155 - Bühler, K., Bühler, B., Klähn, S., Krömer, J.O., Dusny, C., Schmid, A. (2021):
Biocatalytic production of white hydrogen from water using cyanobacteria
In: Rögner, M. (ed.)
Photosynthesis: Biotechnological applications with microalgae
De Gruyter, Berlin ; Boston, p. 279 - 306
10.1515/9783110716979-011 - Bühler, K., Krömer, J.O., Klähn, S., Bühler, B., Dusny, C., Schmid, A. (2021):
Weißer Wasserstoff made in Leipzig [White hydrogen made in Leipzig]
Biospektrum 27 (3), 335
10.1007/s12268-021-1572-x - Klähn, S., Mikkat, S., Riediger, M., Georg, J., Hess, W.R., Hagemann, M. (2021):
Integrative analysis of the salt stress response in cyanobacteria
Biol. Direct 16 , art. 26
10.1186/s13062-021-00316-4 - Lupacchini, S., Appel, J., Stauder, R., Bolay, P., Klähn, S., Lettau, E., Adrian, L., Lauterbach, L., Bühler, B., Schmid, A., Toepel, J. (2021):
Rewiring cyanobacterial photosynthesis by the implementation of an oxygen-tolerant hydrogenase
Metab. Eng. 68 , 199 - 209
10.1016/j.ymben.2021.10.006
2020 (1)
- Brandenburg, F., Klähn, S. (2020):
Small but smart: On the diverse role of small proteins in the regulation of cyanobacterial metabolism
Life 10 (12), art. 322
10.3390/life10120322
2019 (4)
- Gärtner, K., Klähn, S., Watanabe, N., Mikkat, S., Scholz, I., Hess, W.R., Hagemann, M. (2019):
Cytosine N4-methylation via M.Ssp6803II is involved in the regulation of transcription, fine-tuning of DNA replication and DNA repair in the cyanobacterium Synechocystis sp. PCC 6803
Front. Microbiol. 10 , art. 1233
10.3389/fmicb.2019.01233 - Kirsch, F., Klähn, S., Hagemann, M. (2019):
Salt-regulated accumulation of the compatible solutes sucrose and glucosylglycerol in cyanobacteria and its biotechnological potential
Front. Microbiol. 10 , art. 2139
10.3389/fmicb.2019.02139 - Klähn, S. (2019):
Same same but different: Wie Cyanobakterien ein zentrales Enzym regulieren [Same same but different: How cyanobacteria regulate a central enzyme]
Biospektrum 25 (6), 610 - 613
10.1007/s12268-019-0208-x - Klähn, S. (2019):
Mikroorganismus in den Schlagzeilen: Ein natürlicher Glyphosat-Ersatz aus Cyanobakterien?
Biospektrum 25 (5), 536
10.1007/s12268-019-0193-0
2018 (5)
- Bolay, P., Muro-Pastor, M.I., Florencio, F.J., Klähn, S. (2018):
The distinctive regulation of cyanobacterial glutamine synthetase
Life 8 (4), art. 52
10.3390/life8040052 - Hagemann, M., Gärtner, K., Scharnagl, M., Bolay, P., Lott, S.C., Fuss, J., Huettel, B., Reinhardt, R., Klähn, S., Hess, W.R. (2018):
Identification of the DNA methyltransferases establishing themethylome of the cyanobacterium Synechocystis sp. PCC 6803
DNA Res. 25 (4), 343 - 352
10.1093/dnares/dsy006 - Klähn, S., Bolay, P., Wright, P.R., Atilho, R.M., Brewer, K.I., Hagemann, M., Breaker, R.R., Hess, W.R. (2018):
A glutamine riboswitch is a key element for the regulation of glutamine synthetase in cyanobacteria
Nucleic Acids Res. 46 (19), 10082 - 10094
10.1093/nar/gky709 - Rübsam, H., Kirsch, F., Reimann, V., Erban, A., Kopka, J., Hagemann, M., Hess, W.R., Klähn, S. (2018):
The iron-stress activated RNA 1 (IsaR1) coordinates osmotic acclimation and iron starvation responses in the cyanobacterium Synechocystis sp. PCC 6803
Environ. Microbiol. 20 (8), 2757 - 2768
10.1111/1462-2920.14079 - Tan, X., Hou, S., Georg, J., Klähn, S., Lu, X., Hess, W.R. (2018):
The primary transcriptome of the fast-growing cyanobacterium Synechococcus elongatus UTEX 2973
Biotechnol. Biofuels 11 , art. 218
10.1186/s13068-018-1215-8
2017 (3)
- Kuchmina E., Klähn S., Jakob A., Bigott W., Enke H., Dühring U., Wilde A. (2017)
Ethylene production in Synechocystis sp. PCC 6803 promotes phototactic movement.
Microbiology, 163 (12), 1937-1945.
- Kirsch F., Pade N., Klähn S., Hess W.R., Hagemann M. (2017)
The glucosylglycerol-degrading enzyme GghA is involved in acclimation to fluctuating salinities by the cyanobacterium Synechocystis sp. strain PCC 6803.
Microbiology, 163 (9), 1319-1328.
- Georg J., Kostova G., Vuorijoki L., Schön V., Kadowaki T., Huokko T., Baumgartner D., Müller M., Klähn S., Allahverdiyeva Y., Hihara Y., Futschik M.E., Aro E.M., Hess W.R. (2017)
Acclimation of oxygenic photosynthesis to iron starvation is controlled by the sRNA IsaR1.
Current Biology 27 (10):1425-1436.e7.
2016 (3)
- Baumgartner D., Kopf M., Klähn S., Steglich C., Hess WR. (2016)
Small proteins in cyanobacteria provide a paradigm for the functional analysis of the bacterial micro-proteome.
BMC Microbiology 16:285
- Orf I., Schwarz D., Kaplan A., Kopka J., Hess W.R., Hagemann M., Klähn S.* (2016)
CyAbrB2 contributes to the transcriptional regulation of Low CO2 acclimation in Synechocystis sp. PCC 6803.
Plant and Cell Physiology 57(10): 2232–2243.
*corresponding author
- de Porcellinis A.J.*, Klähn S.*, Rosgaard L., Kirsch R., Gutekunst K., Georg J., Hess W.R., Sakuragi Y. (2016)
The non-coding RNA Ncr0700/PmgR1 is required for photomixotrophic growth and the regulation of glycogen accumulation in the cyanobacterium Synechocystis sp. PCC 6803.
Plant and Cell Physiology 57(10): 2091–2103.
*shared first authors
2015 (4)
- Klähn S., Schaal C., Georg J., Baumgartner D., Knippen G., Hagemann M., Muro-Pastor A.M., Hess W.R. (2015)
The sRNA NsiR4 is involved in nitrogen assimilation control in cyanobacteria by targeting glutamine synthetase inactivating factor IF7.
Proceedings of the National Academy of Sciences of the United States of America 112 (45): E6243 - E6252.
- Orf I., Klähn S., Schwarz D., Frank M., Hess W.R., Hagemann M., Kopka J. (2015)
Integrated analysis of engineered carbon limitation in a quadruple CO2/HCO3- uptake mutant of Synechocystis sp. PCC 6803.
Plant Physiology 169: 1787 - 1806
- Kopf M.*, Klähn S.*, Scholz I., Hess W.R., Voß B. (2015)
Variations in the non-coding transcriptome as a driver of inter-strain divergence and physiological adaptation in bacteria.
Scientific Reports 5: 9560.
*shared first authors
- Klähn S., Orf I., Schwarz D., Matthiessen J.K.F., Kopka J., Hess W.R., Hagemann M. (2015)
Integrated transcriptomic and metabolomic characterization of the low-carbon response using an ndhR mutant of Synechocystis sp. PCC 6803.
Plant Physiology 169: 1540 - 1556
2014 (5)
- Georg J., Dienst D., Schürgers N., Wallner T., Kopp D., Stazic D., Kuchmina E., Klähn S., Lokstein H., Hess W.R., Wilde A. (2014)
The small regulatory RNA PsrR1 controls photosynthetic functions in cyanobacteria.
The Plant Cell 26 (9): 3661 - 3679.
- Kopf M., Klähn S., Voß B., Stüber K., Huettel B., Reinhardt R., Hess W.R. (2014)
Finished genome sequence of the unicellular cyanobacterium Synechocystis sp. strain PCC 6714.
Genome Announcements 2 (4): e00757-14.
- Klähn S., Baumgartner D., Pfreundt U., Voigt K., Schön V., Steglich C., Hess W.R. (2014)
Alkane biosynthesis genes in cyanobacteria and their transcriptional organization.
Frontiers in Bioengineering and Biotechnology 2: 24.
- Kopf M.*, Klähn S.*, Scholz I., Matthiessen J.K.F., Hess W.R., Voß B. (2014)
Comparative analysis of the primary transcriptome of Synechocystis sp. PCC 6803.
DNA Research 21 (5): 527 - 539.
*shared first authors
- Kopf M.*, Klähn S.*, Pade N., Weingärtner C., Hagemann M., Voß B., Hess W.R. (2014)
Comparative genome analysis of the closely related Synechocystis strains PCC 6714 and PCC 6803.
DNA Research 21 (3):255 - 266.
*shared first authors, Editor’s choice (Highlighting article in Volume 21, Issue 3)
2013 (1)
- Sievers N., Muders K., Henneberg M., Klähn S., Effmert M., Junghans H., Hagemann M. (2013)
Establishing glucosylglycerol synthesis in potato (Solanum tuberosum L. cv. Albatros) by expression of the ggpPS gene from Azotobacter vinelandii.
Journal of Plant Science and Molecular Breeding 2: 1.
2012 (2)
- Eisenhut M., Georg J., Klähn S., Sakurai I., Mustila H., Zhang P., Hess W.R., Aro E.M. (2012)
The antisense RNA As1_flv4 in the cyanobacterium Synechocystis sp. PCC 6803 prevents premature expression of the flv4-2 operon upon shift in inorganic carbon supply.
Journal of Biological Chemistry 287 (40): 33153 -33162.
- Pade N., Compaoré J., Klähn S., Stal L.J., Hagemann M. (2012)
The marine cyanobacterium Crocosphaera watsonii WH8501 synthesizes the compatible solute trehalose by a laterally acquired OtsAB fusion protein.
Environmental Microbiology 14: 1261 -1271.2009 (2)
2011 (1)
- Klähn S. and Hagemann M. (2011)
Compatible solute biosynthesis in cyanobacteria.
Environmental Microbiology 13: 551 - 562.
2010 (2)
- Klähn S., Höhne A., Simon E., Hagemann M. (2010)
The gene ssl3076 encodes a protein mediating the salt -induced expression of ggpS for the biosynthesis of the compatible solute glucosylglycerol in Synechocystis sp. strain PCC 6803.
Journal of Bacteriology 192: 4403 -4412.
- Klähn S., Steglich C., Hess W.R., Hagemann M. (2010)
Glucosylglycerate: a secondary compatible solute common to marine cyanobacteria from nitrogen-poor environments.
Environmental Microbiology 12: 83 – 94.
2009 (1)
- Klähn S., Marquardt D.M., Rollwitz I., Hagemann M. (2009)
Expression of the ggpPS gene for glucosylglycerol biosynthesis from Azotobacter vinelandii improves the salt tolerance of Arabidopsis thaliana.
Journal of Experimental Botany 60: 1679 - 1689
2008 (1)
- Hagemann M., Ribbeck-Busch K., Klähn S., Hasse D., Steinbruch R., Berg G. (2008)
The plant-associated bacterium Stenotrophomonas rhizophila expresses a new enzyme for the synthesis of the compatible solute glucosylglycerol.
Journal of Bacteriology 190: 5898 - 5906.
Students are always encouraged to join our team. The available projects are interesting for those with a background in microbiology, genetics, molecular biology or related. If you are interested please contact Dr. Stephan Klähn for details.
At the moment for instance, we are seeking for a Master candidate who is helping to design cyanobacterial shuttle vectors. Albeit the biotechnological potential of cyanobacteria is more and more exploited, one of the main problems still compromising their application is the rather limited availability of molecular toolboxes for genetic engineering. Hence, this topic is of general interest for the further development of genetic engineering tools and their use in green biotechnology.
