FATE - Fate and effects of antibiotics from sources to sinks:
Bridging the scale from molecular level processes to large-scale observations in engineered and natural systems
The discovery and use of antibiotics has had a huge impact on human and animal health, but has come at the expense of the rapid rise of antibiotic resistant bacteria. Antibiotic resistance is recognized as one of the major global health threats of this century. Antibiotics are only partially metabolized by humans and animals, and large amounts enter environmental compartments. Antibiotics are also directly released into wastewater from the pharmaceutical industry. A better understanding of the environmental fate and effects of antibiotics is useful to create strategies to improve their degradation and combat antibiotic resistance dissemination.
Knowledge gaps
In 2016, the WHO issued a declaration stating that cross-sectoral cooperation is needed to develop comprehensive and integrative measures to limit the prevalence of clinically relevant antibiotic resistance. This One Health approach recognizes the interlinkage between human, animal and environmental health and emphasizes the importance of close collaboration between all stakeholders. The third interim report of the German Antibiotic Resistance Strategy DART2020 highlighted the need to more strongly integrate the environmental sector into the activities [1].
Recent research on the degradation of antibiotics and the effects on microbial communities in the environment considers the involved compartments separately from each other and does not take into account the mass flow of antibiotics and antibiotic resistant bacteria from source to sink environmental compartments. Currently, there are no standards for the regulation of antibiotics discharge from production industry or other anthropogenic sources [2,3] and also threshold antibiotic concentrations in fertilizer products made from manure or sewage and biogas sludge are not implemented in legislation. So far, only a few antibiotics are included in the ‘watch list’ of the European Water Framework Directive [4]. Determination of acceptable emission levels and environmental quality standards, as well as the development of intervention options for reduced environmental input, requires a thorough understanding of the fate and effects of antibiotics as well as the development, propagation and transmission of antibiotic resistance across the interconnected environmental compartments. FATE aims at addressing these knowledge gaps via the comparative investigation of four environmental compartments representing sources (wastewater treatment and biogas plants) and sinks (agricultural soils and surface waters).
Key Questions
- What is the fate of the target antibiotics in the different environmental compartments?
- How do the target antibiotics contribute to the development and spread of antibiotic resistance?
- What are the effects of the target antibiotics on the indigenous microbial communities?
Research Outline
In the PhD College FATE we want to improve our understanding of the fate and effects of antibiotics and antibiotic resistance along the pathways from environmental source to sink compartments. FATE will make use of state-of-the-art methods and expertise available in the Integrated Projects SusBioTech and CCF and apply them to source (PhD 1 and PhD 2) and sink compartments (PhD 3 and PhD 4) in large-scale experimental facilities and in laboratory experiments. The combined investigation will yield a better understanding of the environmental fate and effects of antibiotics as well as strategies to improve their degradation and combat antibiotic resistance dissemination.
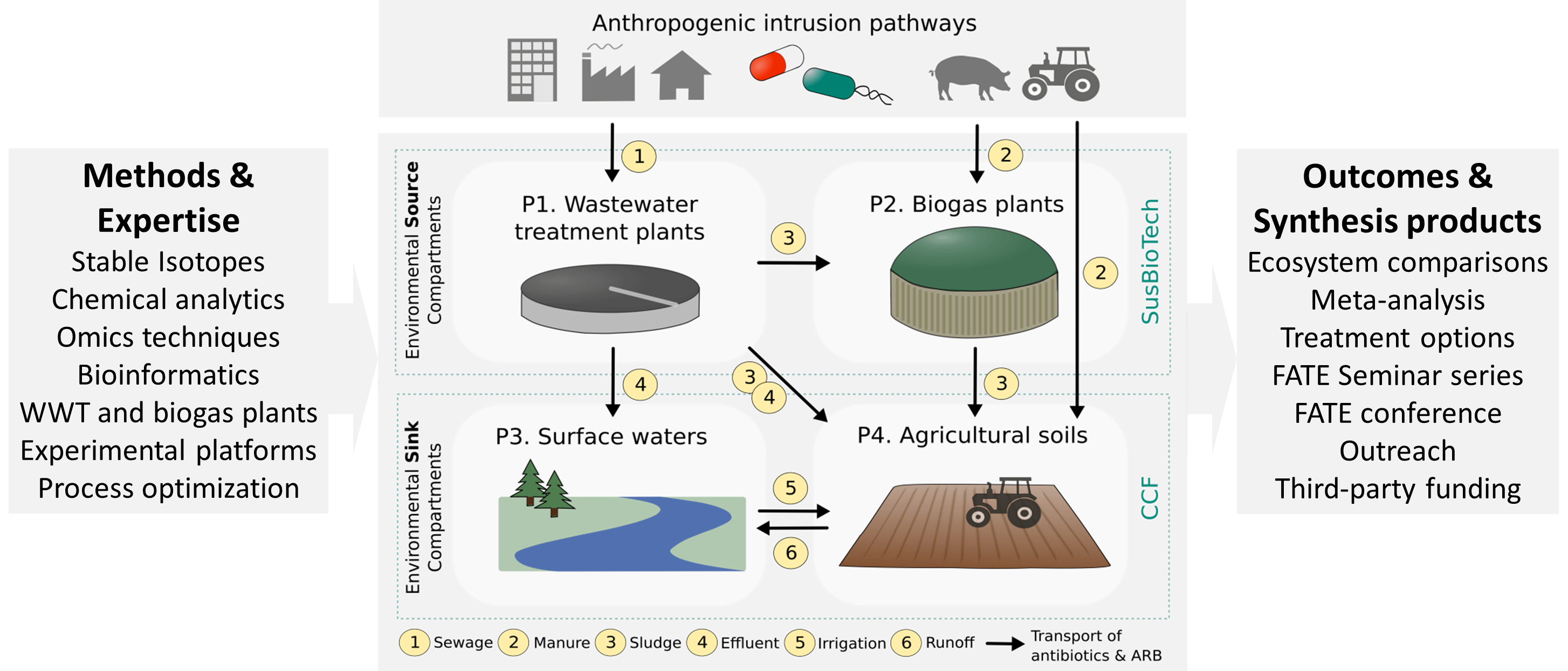
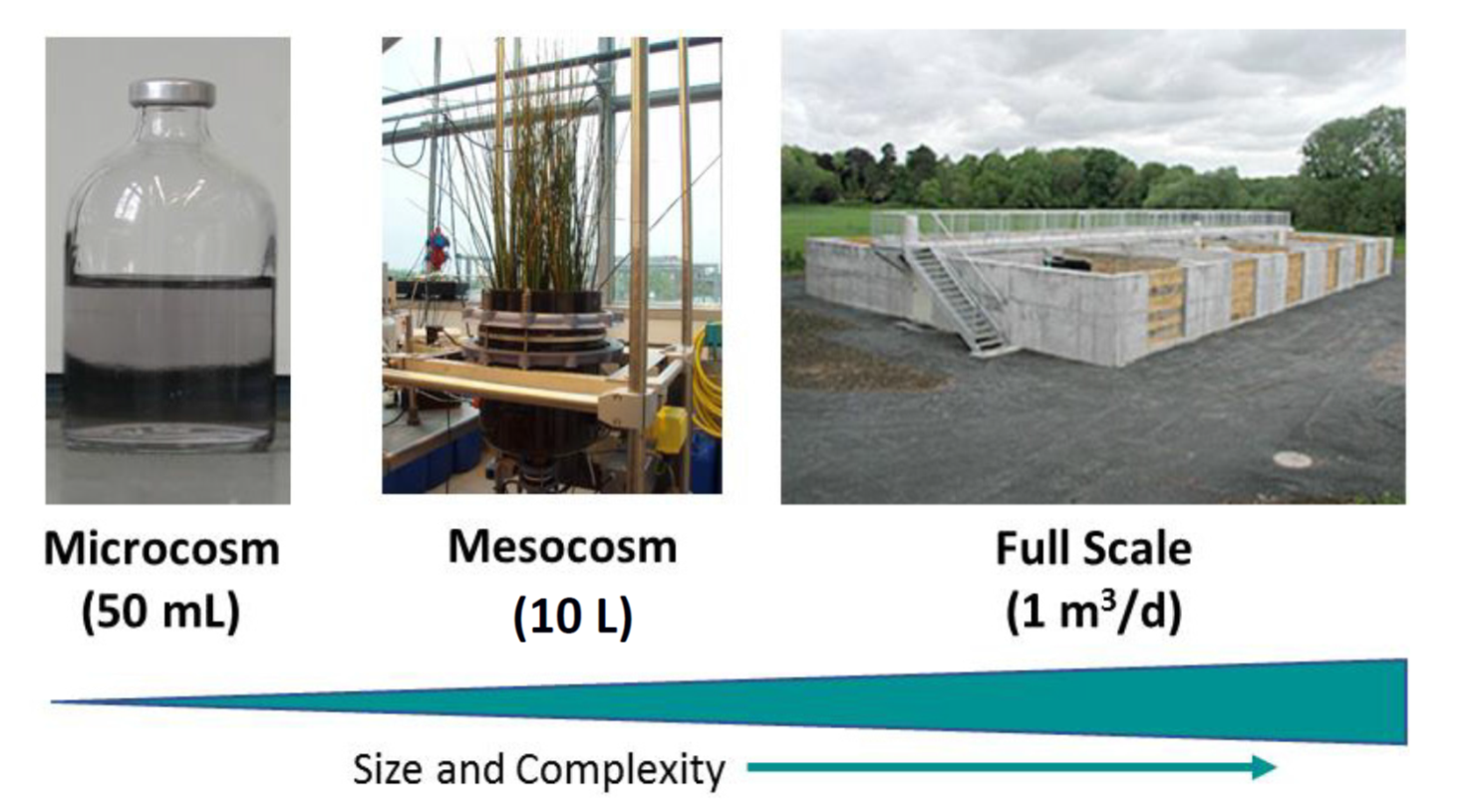
PhD I focus on the anaerobic degradation of residual antibiotics discharged into wastewater. While aerobic degradation of antibiotics is widely studied, it is not known to what extend anoxic processes can contribute to the decrease of their concentrations and which transformation products are generated. This PhD will investigate the interplay of anaerobic, anoxic and oxic transformation of the three target antibiotics sulfamethoxazole, ciprofloxacin, and tetracycline, at different size and complexity scales: lab microcosms (50 mL), mesocosms (10 L), and pilot-scale at the research infrastructure in BDZ.

Key Questions
- Can anoxic pre-treatment decrease antibiotic concentrations in wastewater and which transformation products are formed?
- What is the molecular basis of the catalyzed processes?
- Do anoxic processes already occur in early stages of classical WWT and can knowledge on transformation products foster their detection?
PhD student:
Caglar Akay
Supervising team:
Prof. Dr. Lorenz Adrian (ISOBIO)
Dr. Andrii Butkovskyi (UBT)
Prof. Dr. Roland A. Müller (UBZ)
PhD II will focus on the anaerobic digestion of antibiotics in biogas production. Little is known about the fate of antibiotics and antibiotic resistance genes (ARGs), and the potential horizontal transfer of ARGs during the biogas process. Antibiotic digestion systems constantly receiving loads of antibiotics have the potential of being reservoirs for multi-resistant bacteria. Therefore, a proper management is required to enhance the degradation of antibiotics, while reducing the abundance of ARGs. The goal of this PhD is to transform antibiotic digestion systems, from potential hotspots of antibiotic resistance development and spread, into part of the mitigation strategy.
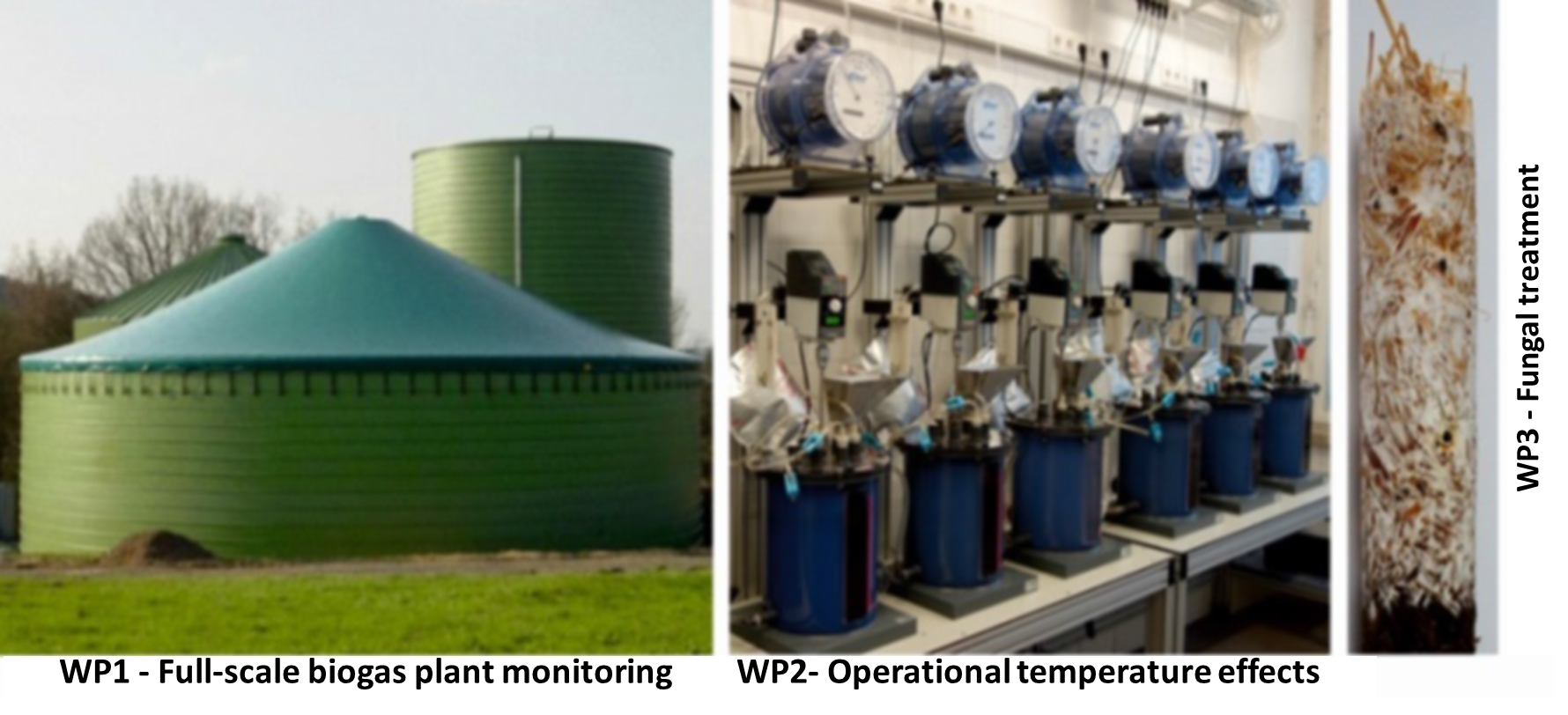

Key Questions
- Can we influence the degradation of antibiotics and removal of ARGs by the management of the reactor operational conditions (e.g. temperature, hydraulic retention time)?
- How do large-scale mesophilic biogas reactors contribute to the degradation of antibiotics and the spread of ARGs?
- Are non-specific enzymes of wood-decaying fungi suitable for the treatment of antibiotics in biogas digestate?
PhD student:
Supervisors:
Dr. Marcell Nikolausz (UMB)
Dr. Hans Hermann Richnow (ISOBIO)
Dr. Jochen Müller (UBT)
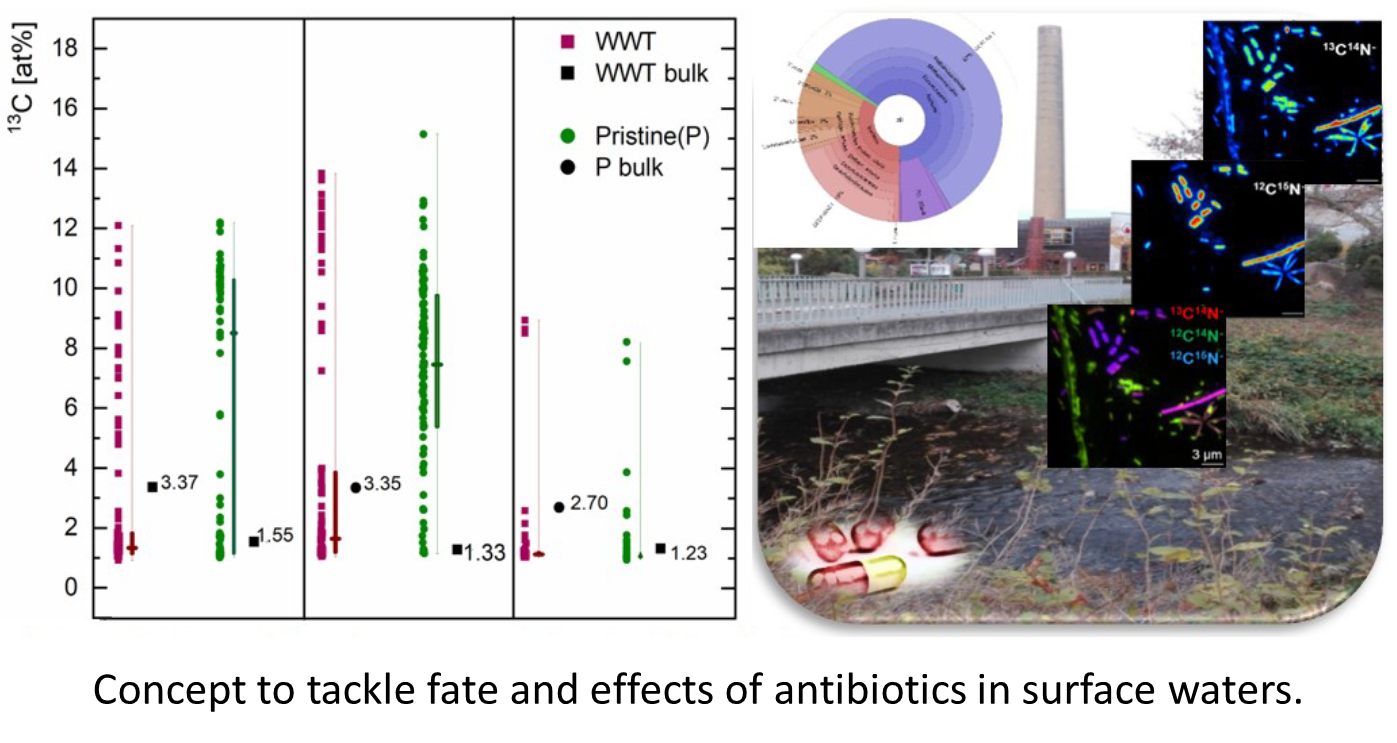
Key Questions
- To what extent do antibiotics impact microbial activity and diversity in natural pristine and wastewater-impacted surface waters?
- Does antibiotic-impacted microbial activity and diversity correlate with the occurrence and spread of antibiotic resistance genes in natural communities?
- What are the primary mechanisms of antibiotic transformation in surface waters and how is this related among scales?
PhD student:
Supervisors:
Dr. Niculina Musat (ISOBIO)
Dr. Hans Hermann Richnow (ISOBIO)
Dr. Jochen Müller (UBT)
PhD IV will focus on the fate and effects of antibiotics in agricultural soils. Once added to the soil, antibiotics interact with soil particles, are subject to microbial transformation, and influence the indigenous microbial community. This PhD will combine literature synthesis, greenhouse and lab-scale experiments to explore the impact of manure fertilization on the soil resistome and to study the particular role of the rhizosphere for antibiotic degradation and ARG development and spread.
Key Questions
- Does manure application affect the abundance and diversity of ARGs in soils?
- Does the rhizosphere boost microbial degradation of antibiotics?
- Do plant endophytes contribute to the degradation of antibiotics?
PhD student:
please apply hereSupervising team:
Dr. Anja Worrich (UMB)Dr. Niculina Musat (ISOBIO)
Dr. Ulisses Nunes da Rocha (UMB)
Dr. Hans Hermann Richnow (ISOBIO)
- Abu Sin, M., et al., Globale und nationale Strategien gegen Antibiotikaresistenzen. Bundesgesundheitsblatt - Gesundheitsforschung - Gesundheitsschutz, 2018. 61(5): p. 507-514.
- Bengtsson-Palme, J., L. Gunnarsson, and D.J. Larsson, Can branding and price of pharmaceuticals guide informed choices towards improved pollution control during manufacturing? Journal of cleaner production, 2018. 171: p. 137-146.
- Bengtsson-Palme, J., E. Kristiansson, and D.J. Larsson, Environmental factors influencing the development and spread of antibiotic resistance. FEMS microbiology reviews, 2017. 42(1): p. fux053.
- Carvalho, R.N., et al., Development of the first watch list under the environmental quality standards directive. JRC Science Hub, 2015.




