Dr. Ulisses Nunes da Rocha
Contact
Dr. Ulisses Nunes da Rocha
Senior Scientist
Microbial Data Science group leader
ulisses.rocha@ufz.de / Phone +49 341 235-1374

Department of Computational Biology and Chemistry
Helmholtz Centre for Environmental Research - UFZ
Permoserstr. 15, 04318 Leipzig, Germany
In my current role as group leader of the Microbial Data Science group at the Helmholtz Centre for Environmental Research – UFZ in Leipzig, I lead a team of 1 postdoctoral researcher, 6 PhD students, 3 MSc students and visiting scholars. Together, we integrate multi-omics data, machine learning and ecological theory to build “digital twins” of microbial systems in engineered and natural environments — enabling the anticipation of community stability, resilience and ecosystem function.
By merging environmental microbiology with scalable computational workflows, I aim to translate microbial complexity into actionable scientific rationale for ecosystem protection and design.
Scientific Career
07/2025 Present
Senior Scientist - Microbial Systems Data Science group, Department Computational Biology and Chemistry, Helmholtz Centre for Environmental Research - UFZ, Leipzig, Germany
04/2021 06/2025
Senior Scientist - Microbial Systems Data Science group, Department of Environmental Microbiology, Helmholtz Centre for Environmental Research - UFZ, Leipzig, Germany
03/2019 present
Guest Professor - Theoretical Microbial Ecology, Institute of Mathematics and Computer Sciences (ICMC), University of Sao Paulo (USP), Sao Carlos, SP, Brazil.
03/2017 03/2021
Junior Group Leader - Microbial Systems Data Science group, Helmholtz Young Investigator, Department of Environmental Microbiology, Helmholtz Centre for Environmental Research - UFZ, Leipzig, Germany
09/2013 11/2016
Post-Doctoral Associate - VU University of Amsterdam, Faculty of Earth and Life Sciences, Department of Molecular Cell Physiology, Amsterdam, the Netherlands
07/2011 08/2013
Post-Doctoral Associate - Emerging Pathogens Institute, University of Florida, Gainesville, FL, USA
02/2005 02/2006
Education
PhD
2006
2010
Microbial
Ecology
University of Groningen, Groningen, The Netherlands
MSc
2003
2005
Agricultural Microbiology
1999
2002
Life Sciences
As an environmental microbiologist, I develop ecological theories and models that embrace prediction and control of functions performed by complex microbial communities. Broad-scale ecosystem functioning is often an intricate interplay between multiple interdependent species and abiotic factors. The complexity behind these systems, ranging from hundreds to thousands of species, makes it immensely difficult to identify the role of all key players involved and to gain a full understanding of ecosystem functioning. With my team, I have attempted to diminish the boundaries between natural sciences and engineering with focus on complex microbial communities. To this end, I use state of the art methods and concepts used in Community Systems Biology:
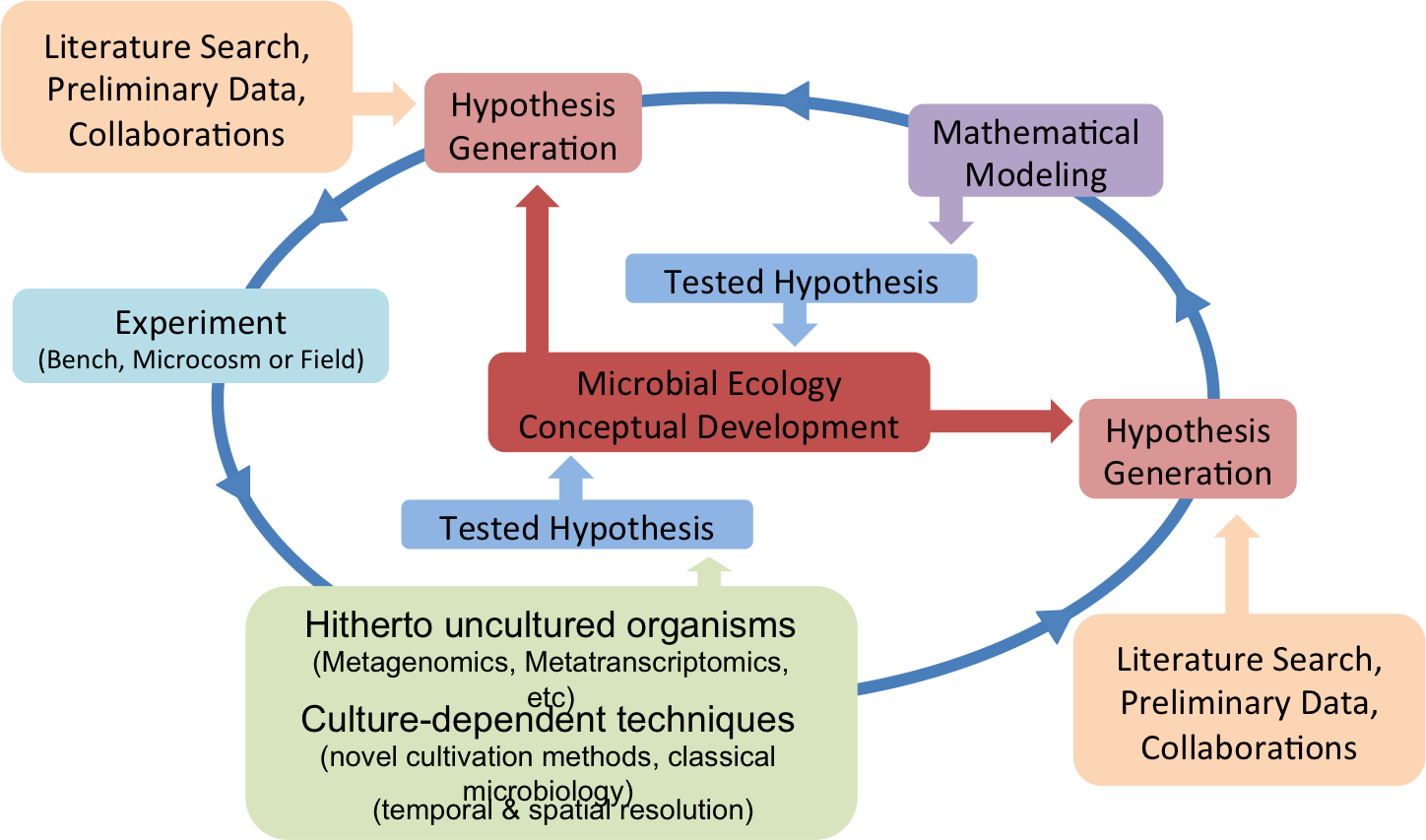
My circular approach allows the study of biochemical limitations of the community and their environmental constraints. Different substrate availability scenarios are modeled and tested to discover how environmental disturbances affect ecosystem functioning. I believe a circular frame between hypothesis testing and conceptual development is key to expose potential resilience of ecosystems to the disturbances and lead to better environmental risk assessment. I am a proven instrumental player linking wet lab and field data to theoretical microbial ecology and computational sciences.
Vision:
To create an interdisciplinary systems biology approach where Environmental Microbiology, Modeling, Microbial Physiology, Bioinformatics and Engineering meet to create out-of-the-box thinking towards prediction and control of microbial interactions in complex microbial communities.
Mission:
Environmental health regulates the conditions, under which life is
sustained on Earth. A clear definition of environmental health is
necessary to allow the implementation of practices that guarantee
sustainable exploration of our limited natural resources. To diminish the boundaries between natural sciences and engineering, I will work to define environmental
health by exploring microbial interactions. I want to achieve this
mission by developing concepts and theories to expand modeling of
microbial communities to the vast diversity found in natural ecosystems
with focus on prediction and control of ecosystem processes during
disturbance events.
Based on my mission and vision, my hypothesis-driven systems approach is grounded on three main research areas:
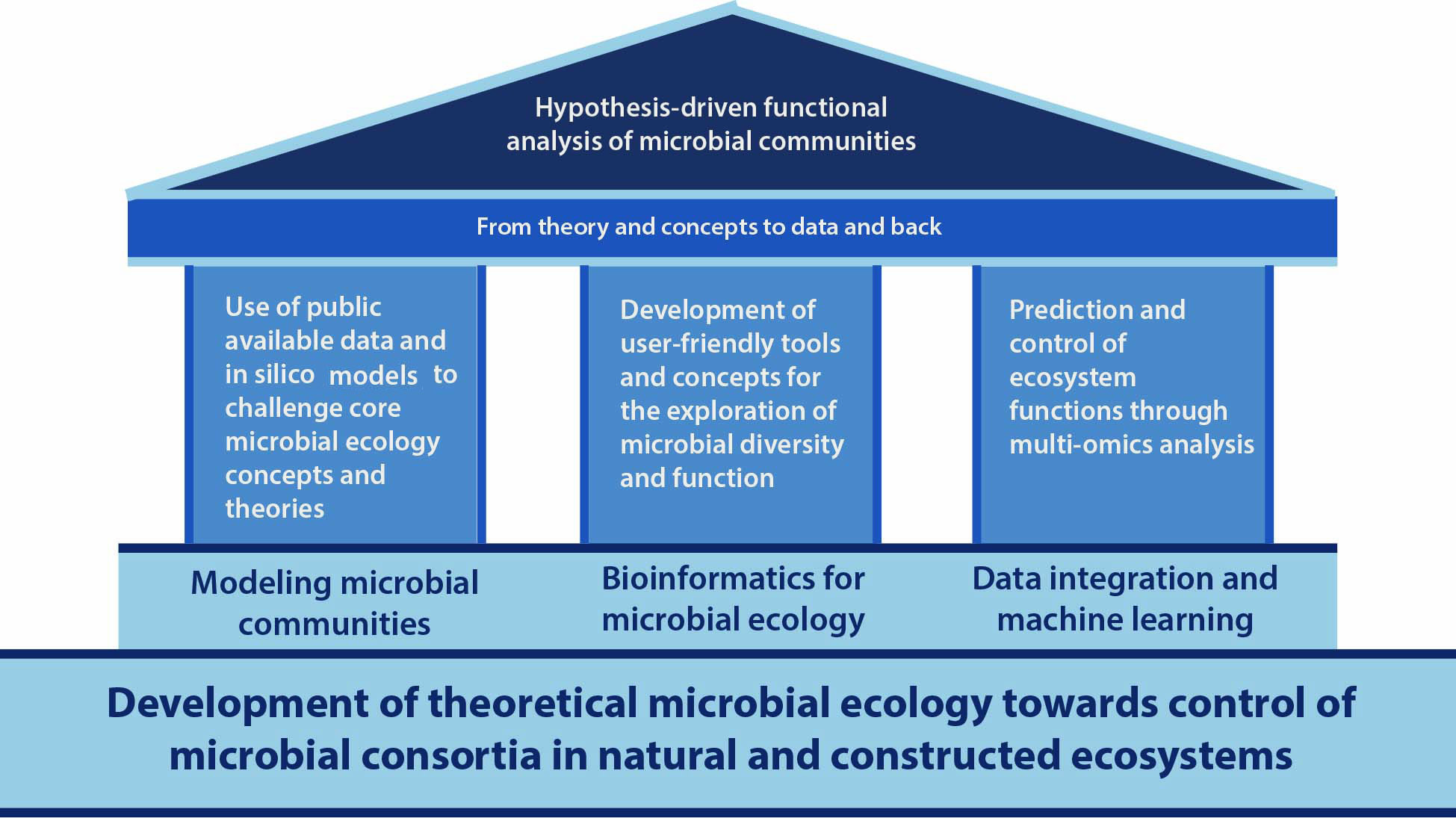
updated on 12-May-2020
- Development of user-friendly bioinformatics tools for the analysis of complex microbial communities. Collaboration between my team at the Helmholtz Centre for Environmental Research (Leipzig, Germany) and Prof. Dr. Peter Stadler at the University of Leipzig. September 2017 – Present. Prof. Stadler and I supervise 2 PhD students. Currently, we are developing web applications to help non-specialists to analyze omics and multi-omics data. In our first web application our team organized metadata of over 28 thousand metagenomes that can now easily be screened and added to metastudies. This web application can be found in the following link: https://webapp.ufz.de/tmdb/
- Detection of dysbiosis bioindicators in human microbiomes via machine learning. Collaboration between my team at the Helmholtz Centre for Environmental Research (Leipzig, Germany) and Prof. Dr. André Carlos Ferreira de Carvalho at University of São Paulo (São Carlos, Brazil). October 2018 This cooperation is based on a double degree program where we will develop concepts and theories to advance the knowledge of prediction of dysbiosis within the field of Theoretical Microbial Ecology of human microbiomes.
- Prediction of ecosystem processes using machine learning in environmental microbial communities (bioreactor and soils) with different levels of complexity. Collaboration between my team at the Helmholtz Centre for Environmental Research (Leipzig, Germany) and Prof. Dr. João Carlos Setubal at University of São Paulo (São Paulo, Brazil). Nov 2018 – Present.
- Study of molecular and ecological mechanisms involved with distribution patterns of marine microbes and host interactions towards microbial holobiome. Collaboration between my team at the Helmholtz Centre for Environmental Research (Leipzig, Germany) and Prof. Dr. Michael Sweet at Derby University (Derby, England). July 2019 – Present. Initial stages of collaboration (preparing marine samples for amplicon sequencing and planning experimental design for multi-omics studies).
- Development of concepts and theories to estimation of regulatory networks in microbial communities through prediction of Transcription Factors. Collaboration between my team at the Helmholtz Centre for Environmental Research (Leipzig, Germany) and Prof. Dr. Rafael Silva-Rocha at University of Sao Paulo (Ribeirão Preto, Brazil). Mar 2018 – Present.
- Prospection of Antarctic Volcanoes microbiomes towards biotechnological applications. Collaboration between my team at the Helmholtz Centre for Environmental Research (Leipzig, Germany) and Prof. Dr. Alexandre Rosado at the Federal University of Rio de Janeiro (Rio de Janeiro, Brazil). Mar 2018 – Present.
- Prediction of Bioindicators using machine learning and recovery of metagenome assembled genomes (Eukaryotes, Prokaryotes and viruses) in deep marine samples for biotechnological applications. Collaboration between my team at the Helmholtz Centre for Environmental Research (Leipzig, Germany) and Prof. Dr. Vivian Pellizari at Sao Paulo University (São Paulo, Brazil). Oct 2018 – Present.
- Study of molecular and ecological mechanisms involved with cycle of organic matter in forest soil. Collaboration between my team at the Helmholtz Centre for Environmental Research (Leipzig, Germany) and Dr. Petr Baldrian at the Czech Academy of Science (Prague, Czech Republic). Jan 2018 – Present.
- Prediction of Bioindicators using machine learning and recovery of metagenome assembled genomes (Eukaryotes, Prokaryotes and viruses) in marine sponges and aquaculture. Collaboration between my team at Helmholtz Centre for Environmental Research and Dr. Newton Gomes at the University of Aveiro (Portugal). Aug 2017 - Present.
- Prediction of Bioindicators using machine learning and recovery of metagenomic assembled genomes (Eukaryotes, Prokaryotes and viruses) in corals. Collaboration between my team at Helmholtz Centre for Environmental Research and Prof. Dr. Raquel Peixoto at the Federal University of Rio de Janeiro. Jun 2017 - Present.
- Study of the bacterial diversity in marine sponges. Collaboration between my team at Helmholtz Centre for Environmental Research and Prof. Dr. Rodrigo Costa at the Technical University of Lisboa. Jun 2017 - Present.
updated 01-Oct-2019
- Helmholtz Young Investigator Grant (2017-2023): Project Tittle: ‘Micro ‘Big Data’: Mining Microbial Functional Hotspots in Terrestrial Environments’. Budget: 1.8 Million Euros.
- Double degree program University of Sao Paulo (SP, Brazil) and Microbial Systems Data Science Group (UFZ) (2019-2023). 2 PhD students financed to stay 2 years in Brazil and 1.5 years in Germany. PhD 1 project tittle: ‘Detection of dysbiosis bioindicators in human microbiomes via machine learning’. PhD 2 project tittle: ‘Predicting carbon, nitrogen and phosphorus cycles in soils from microbiome data using machine learning’.
- BfR Biorezistans (2019-2020) – Search for mechanisms of antimicrobial resistance in Listeria monocytogenes, a multi-omics approach. (BfR grant, 80 thousand Euros).
- Portuguese Foundation for Science & Technology (FCT). Project Title: Harnessing the power of the microbial metamobilome: using marine sponges as models to uncover novel biotechnological appliances from symbiont communities. Principal Investgator: Rodrigo da Silva Costa (2016-2022). 190 Thousand Euros.
- Portuguese Foundation for Science & Technology (FCT). Project Title: Illuminating the metabolism of foundational and elusive microbial symbionts in benthic ecosystems: a cultivation-independent quest for Hahellaceae (Gammaproteobacteria, Oceanospirillales) genomes. Principal Investigator: Rodrigo da Silva Costa (2014-2015). 50 Thousand Euros.
- Portuguese Foundation for Science & Technology (FCT). Project Title: Single cell genomics and metagenomics-aided cultivation of the unculturable: uncovering the life-style of Acidobacteria symbionts of marine sponges. Principal Investigator: Rodrigo da Silva Costa (2013-2015). 110 Thousand Euros.
- Community Sequencing Program. Project Title: Mediterranean Grassland Soil Metagenome (MGSM): Enabling a systems view of soil carbon and nitrogen biogeochemistry under a changing climate. Principal Investigator: Eoin Brodie (2012-2013). 170 thousand dollars.
Publications
Summary:
Total number of citations: 7,319; H-index: 32 (both numbers collected from Scopus on November 20, 2025).
Authors followed by a # contributed equally to the publication / IF – Impact factor of journals
H8. [+3] and da Rocha UN (2020). TerrestrialMetagenomeDB: a public repository of curated and standardized metadata for terrestrial metagenomes. Nucleic Acids Research, 48(D1) (IF: 11.147).H7. [+5], da Rocha UN, [+5] (2019). Marine probiotics: increasing coral resistance to bleaching through microbiome manipulation. ISME Journal, 13: 921-936 (IF: 9.493)
H6. [+2], da Rocha UN, [+7] (2019). Microbial community drivers of natural product genediversity in selected soils. Microbiome, 7:78 (IF: 10.465)
H5. [+4], da Rocha UN, [+8] (2018). Dynamic root exudate chemistry and microbial substrate preferences drive patterns in rhizosphere microbial community assembly. Nat Microbiol, 3:470-480. (IF: 14.300)
H4. [+3], da Rocha UN, [+3] (2016). Bacteria increase arid land soil surface temperature through the production of sunscreens. Nature Communications, 7:10373. (IF: 11.878)
H3. Pylro VS, Mui TS, Gilbert J, …, da Rocha UN, Sarmento H, Roesch LFW (2016). A step forward to empower global microbiome research through local leadership. Trends Microbiol, 24: 767-771. (IF: 11.974)
H2. [+2], da Rocha UN, [+6] (2015). Exometabolomics reveals diverse substrate preferences among soil bacteria. Nature Communications, 6:8289. (IF: 11.878)
H1. da Rocha UN#, Rajeev L#, Klitgord N#, [+11] (2013). Dynamic cyanobacterial response to hydration and dehydration in a desert biological soil crust. ISME J, 77: 2178-2191. (IF: 9.493)
This list excludes accepted and submitted publications
Authors followed by a # contributed equally to the publication
IF – Impact factor of journals
For up to date list check https://orcid.org/0000-0001-6972-6692
- 43. Melkonian C, Fillinger F, Atashgahi S, da Rocha UN, Kuiper, E, Olivier B, Braster M, Gottstein W, Helmus R, Parson J, Smidt H, van de Walls M, Gerritse J, Brandt BW, Molenaar D, van Spanning R (2021). Biodiversity and niche partitioning in an anaerobic benzene degrading culture. Communications Biology, Accepted.
- 42. Keller-Costa T, Lago-Lestón A, Saraiva JP, Toscan RB, Silva SG, Gonçalves J, Cox CJ, Kyrpides N, da Rocha UN, Costa R (2021). Metagenomic insights into the taxonomy, function and dysbiosis of prokaryotic communities in octocorals. Microbiome, Accepted (IF: 11.607)
- 41. Tláskal V, Brabcová V, Větrovský T, Jomura M, López-Mondéjar R, Oliveira Monteiro LM, Saraiva JP, Rainier Human Z, Cajthaml T, da Rocha UN, Baldrian P. (2021) Complementary Roles of Wood-Inhabiting Fungi and Bacteria Facilitate Deadwood Decomposition. mSystems 6:1, e01078-20 (IF: 6.280)
- 40. Metze D, Popp D, Schwab L, Keller N-S, da Rocha UN, Richnow H-H, Vogt C (2021). Temperature management potentially affects carbon mineralization capacity and microbial community composition of a shallow aquifer. FEMS Microbiol Ecol 97:2, fiaa261 (IF: 3.675)
- 39. Kasmanas JC, Bartholomäus A, Corrêa FB, Tal T, Jehmlich N, Herberth G, von Bergen M, Stadler PF, de Leon Ferreira de Carvalho ACP, da Rocha UN (2021). HumanMetagenomeDB: A public repository of curated and standardized metadata for human metagenomes. Nucleic Acid Res 49, D743-D750 (IF: 11.501)
- 38. Oliveira V, Polónia ARM, Cleary DFR, Huang YM, de Voogd NJ, da Rocha UN, Gomes NCM (2021). Characterization of putative circular plasmids in sponge-associated bacterial communities using a selective multiply-primed rolling circle amplification. Mol Ecol Resour 21: 1, 110-121 (IF: 7.059)
- 37. Lian S, Nikolausz M, Nijenhuis I, da Rocha UN, Liu B, Corrêa FB, Saraiva JP, Richnow HH (2020). Biotransformation of hexachlorocyclohexanes contaminated biomass for energetic utilization demonstrated in continuous anaerobic digestion system. J Hazard Mater 384, 121448 (IF: 9.038)
- 36. Feng L, Perschke YML, Fontaine D, Nikolausz M, Ward AJ, da Rocha UN, Corrêa FB, Eriksen J, Sørensen P, Møller HB (2020). Anaerobic digestion of co-ensiled cover crop and barley straw: Effect of co-ensiling ratios, manure addition and impact on microbial community structure. Ind Crops Prod 144, 112025 (IF: 4.244)
- 35. López-Mondéjar R, Tláskal V, Větrovský T, Štursová M, Toscan R, da Rocha UN, Baldrian P (2020). Metagenomics and stable isotope probing reveal the complementary contribution of fungal and bacterial communities in the recycling of dead biomass in forest soil. Soil Biol Biochem 148, 107875 (IF: 5.795)
- 34. Bernardes JP, ... da Rocha UN,..., Rosenstiel P (2020). Longitudinal Multi-omics Analyses Identify Responses of Megakaryocytes, Erythroid Cells, and Plasmablasts as Hallmarks of Severe COVID-19. Immunity 53: 6, 1296-1314.e9 (IF: 22.553)
- 33. Schulte-Schrepping J, ... da Rocha UN,..., Schultze JL (2020). Severe COVID-19 Is Marked by a Dysregulated Myeloid Cell Compartment. Cell 182: 6, 1419-1440.e23 (IF: 38.637)
- 32. Marques M, Borges N, Silva SG, da Rocha UN, Lago-Lestón A, Keller-Costa T, Costa R (2020). Metagenome-assembled genome sequences of three uncultured Planktomarina sp. Strains from the Northeast Atlantic Ocean. Microbiol Resour Announc 9:12, e00127-20 (IF: 0.413)
- 31. Corrêa FB, Saraiva JP, Stadler PF and da Rocha UN (2020). TerrestrialMetagenomeDB: a public repository of curated and standardized metadata for terrestrial metagenomes. Nucleic Acid Res 48, D626–D632 (IF: 11.147)
- 30. Liana S, Nikolausz M, Nijenhuis I, da Rocha UN, Liu B, Borim Corrê F, Saraiva JP, Richnow HH (2019). Biotransformation of hexachlorocyclohexanes contaminated biomass for energetic utilization demonstrated in continuous anaerobic digestion system. J Harzard Mater 384, 121448 (IF: 7.650)
- 29. Schultz J, Kallies K, da Rocha UN, Rosado AS (2019). Draft-genome Sequence of Geobacillus sp. strain LEMMJ02, a thermophile isolated from an Antarctic volcano. Microbiol Res Announ 8:e00920-19. (IF: 0.486)
- 28. Schultz J, Kallies R, da Rocha UN,Rosado AS (2019). Draft genome sequence of Brevibacillus sp. strain LEMMJ03, isolated froman Antarctic volcano. Microbiol Resour Announc 8:e00921-19. (IF: 0.486)
- 27. Kallies R, Toscan RB, da Rocha UN, Anders J, Marz M, Chatzinotas A (2019). Evaluation of sequencing library preparation protocols for viral metagenomic analysis from pristine aquifer groundwaters. Viruses, 11(6), 484. (IF: 3.811)
- 26. Borsetto C, Amos GCA, da Rocha UN, Mitchel AL, Finn RD, Laidi RF, Vallin C, Pearce DA, Newsham KK, Wellington EMH (2019). Microbial community drivers of natural product genediversity in selected soils. Microbiome, 7:78 (IF: 10.465)
- 25. Karimi E, Keller-Costa T, Slaby BM, Cox CJ, da Rocha UN, Hentschel U, Costa R (2019). Genomic blueprints of sponge-prokaryote symbiosis are shared by low abundant and cultivatable Alphaproteobacteria. Scientific Reports, volume 9, Article number: 1999. (IF: 4.525)
- 24. Rosado PM, Leite DCA, Duarte GAS, Chaloub RM, Jospin G, da Rocha UN, Saraiva JP, Dini-Andreote F, Eisen JA, Bourne DG & Peixoto RS (2019). Marine probiotics: increasing coral resistance to bleaching through microbiome manipulation. ISME Journal, 13: 921-936 (IF: 9.493)
- 23. Zhalnina K, Louie KB, Hao Z, Mansoori N, da Rocha UN, Shi S, Cho H, Karaoz U, Loqué D, Bowen BP, Firestone MK, Northen TR, Brodie EL (2018). Dynamic root exudate chemistry and microbial substrate preferences drive patterns in rhizosphere microbial community assembly. Nat Microbiol, 3:470-480. (IF: 14.300)
- 22. Karaoz U, Couradeau E, da Rocha UN, Lima H-C, Northen T, Garcia-Pichel F, Brodie EL (2018). Large Blooms of Bacillales (Firmicutes) Underlie the Response to Wetting of Cyanobacterial Biocrusts at Various Stages of Maturity. mBio, 9: e01366-16. (IF: 6.747)
- 21. Atashgahi S, Hornung B, Van Der Waals MJ, da Rocha UN, Hugenholtz F, Nijsse B, Molenaar D, Van Spanning R, Stams AJM, Gerritse J, Smidt H (2018). A benzene-degrading nitrate-reducing microbial consortium displays aerobic and anaerobic benzene degradation pathways. Sci Rep, 8:4490 (IF: 4.525)
- 20. Van der Walls M, Atashgahi S, da Rocha UN, Van der Zaan BM, Smidt H, Gerritse J (2017). Benzene degradation in a denitrifying biofilm reactor: activity and microbial community composition. Appl Microbiol Biotechnol, 101:5175-5188 (IF: 3.670).
- 19. Pylro VS, Mui TS, Gilbert J, …, da Rocha UN, Sarmento H, Roesch LFW (2016). A step forward to empower global microbiome research through local leadership. Trends Microbiol, 24: 767-771. (IF: 11.974)
- 18. Couradeau E, Karaoz U, Lim HC, da Rocha UN, Northern TR, Brodie EL, and Garcia-Pichel F (2016). Bacteria increase arid land soil surface temperature through the production of sunscreens. Nature Communications, 7:10373. (IF: 14.300)
- 17. Shi S, Nuccio E, Herman DJ, Rijkers R, Estera K, Li J, da Rocha UN, He Z, Pett-Ridge J, Brodie EL, Zhou J, Firestone M (2015). Successional trajectories of rhizosphere bacterial communities over consecutive seasons. mBio, 6(4):e00746-15, 8p. (IF: 6.747)
- 16. da Rocha UN, Cadillo-Quiroz H, Karaoz U, Rajeev L, Klitgord N, Dunn S, Truong V, Buenrostro M, Bowen BP, Garcia-Pichel F, Mukhopadhyay A, Northen TR, and Brodie EL (2015). Isolation of a significant fraction of non-phototroph diversity from a desert Biological Soil Crust. Front Microbiol 6:277. (IF: 4.259)
- 15. Baran R, Mayberry-Lewis J, da Rocha UN, Bowen BP, Karaoz U, Brodie EL, Cadillo-Quiroz H, Garcia-Pichel F, and Northen TR (2015). Exometabolomics reveals diverse substrate preferences among soil bacteria. Nature Communications, 6:8289. (IF: 11.878)
- 14. da Rocha UN, Plugge C, George I, van Elsas JD, and van Overbeek LS (2013). The rhizosphere is a habitat that selects for particular groups of Acidobacteria and Verrucomicrobia. PlosOne, e82443. (IF: 2.776)
- 13. Shen W, Cevallos-Cevallos JM, da Rocha UN, Arevalo HA, Stansly PA, Roberts PD, and van Bruggen AHC (2013). Relation between plant nutrition, hormones, insecticide applications, bacterial endophytes and Huanglongbing in citrus trees. Eur J Plant Pathol, 137: 727-742. (IF: 1.744)
- 12. da Rocha UN#, Rajeev L#, Klitgord N#, Luning EG, Fortney J, Axen SP, Shih PM, Bouskill NJ, Bowen BP, Kerfield C, Garcia-Pichel F, Brodie EL, Northen TR, and Mukhopadhyay A (2013). Dynamic cyanobacterial response to hydration and dehydration in a desert biological soil crust. ISME J, 77: 2178-2191. (IF: 9.493)
- 11. Costa R, Keller-Costa T, Gomes NCM, da Rocha UN, van Overbeek L, and van Elsas JD (2013). Evidence for selective bacterial community structuring in the freshwater sponge Ephydatia fluviatilis. Microb Ecol, 65: 232-244. (IF: 3.611)
- 10. da Rocha UN, van Elsas JD, and van Overbeek L (2011). Verrucomicrobia subdivision 1 strains display a difference in the colonization of the leek (Allium porrum) rhizosphere. FEMS Microbiol Ecol, 78: 297-305. (IF: 4.098)
- 9. van Overbeek LS, Franke AC, Nijhuis EHM, Groeneveld RMW, da Rocha UN, and Lotz LAP (2011). Bacterial community structures of Chenopodium album and Stellaria media seeds from arable soils. Microb Ecol, 62: 257-264. (IF: 3.611)
- 8. da Rocha UN, van Elsas JD, and van Overbeek LS (2010). Real-time PCR detection of Holophagae (Acidobacteria) and Verrucomicrobia subdivision 1 groups in bulk and leek (Allium porrum) rhizosphere soils. J Microbiol Methods, 83: 141-148. (IF:1.803)
- 7. Andreote FD, da Rocha UN, Araújo WL, Azevedo JL, and Van Overbeek LS (2010). Effect of bacterial inoculation, plant genotype and developmental stage on root associated and endophytic bacterial communities in potato (Solanum tuberosum). Antonie van Leeuwenhoek, 97: 389-399. (IF: 1.934)
- 6. da Rocha UN, Andreoti FD, Azevedo JL, van Elsas JD, and van Overbeek L (2010). Cultivation of hitherto uncultured bacteria belonging to the Verrucomicrobia subdivision 1 from the potato (Solanum tuberosum) rhizosphere. J Soil Sediments, 10: 326-33. (IF: 2.669)
- 5. da Rocha UN, van Overbeek L, van Elsas JD (2009). Exploration of hitherto-uncultured bacteria from the rhizosphere. FEMS Microbiol Ecol, 69: 3396-3406. (IF: 4.098)
- 4. Andreote FD, De Araújo WL, De Azevedo JL, van Elsas JD, da Rocha UN, and van Overbeek LS (2009). Endophytic colonization of potato (Solanum tuberosum L.) by a novel competent bacterial endophyte, Pseudomonas putida strain P9, and its effect on associated bacterial communities. Appl Environm Microbiol, 75: 3396-3406. (IF: 4.077)
- 3. da Rocha UN, Tótola MR, Pessoa DMM, Júnior JTA, Neves JCL, and Borges AC (2009). Mobilisation of bacteria in a fine-grained residual soil by electrophoresis. J Hazard Mat, 161: 485-491. (IF: 9.063)
- 2. van Overbeek L, Gassner F, van der Plas CL, Kastelein P, da Rocha UN, and Takken W (2008). Diversity of Ixodes ricinus tick-associated bacterial communities from different forests. FEMS Microbiol Ecol, 66: 77-84. (IF: 4.098)
- 1. Borges MT, Nascimento AG, da Rocha UN, and Tótola MR (2008). Nitrogen starvation affects bacterial adhesion to soil. Braz J Microbiol, 39: 457-463. (IF: 2.857)
- 4. da Rocha UN (2019). Isolation of Uncultured Bacteria. In Modern Soil Microbiology, third edition, CRC Press, J.D. van Elsas (chief Editor), ISBN 9781498763530.
- 3. Costa R, Kimes NE, da Rocha UN, López-Pérez M, Esteves AIS, Mizuno C (2016). Marine Microbial Diversity. In Molecular Diversity of Environmental Prokayotes, CRC Press, A.E. Trindade Silva (chief Editor). ISBN 9781482233223.
- 2. Röling WFM, Fillinger L, and da Rocha UN (2014). Analysis of the hierarchical and metabolic regulation of flux through metabolic pathways. In Hydrocarbon and Lipid Microbiology Protocols, Springer Protocols Handbooks,T.J. McGenity et al. (eds.), DOI 10.1007/8623_2014_6, © Springer-Verlag Berlin Heidelberg 2014.
- 1. Röling WFM, Fillinger L, and da Rocha UN (2014). Analysis of the regulation of the rate of hydrocarbon and nutrient flow through microbial communities. In Hydrocarbon and Lipid Microbiology Protocols, Springer Protocols Handbooks,T.J. McGenity et al. (eds.), DOI 10.1007/8623_2014_15, © Springer-Verlag Berlin Heidelberg 2014.
1. Araruna Jr JT, Pessoa DMM, Mergulhão AJPP, Souza MV, da Rocha UN, and Nacimento AR. Nutrient transport and microbial stimulation in residual soil by electrokinetics. In Situ and On-Site Bioremediation Jun 2003 Orlando - Florida. Anals of 7th International In Situ and On-Site Bioremediation Symposium. Columbus - OH, USA: Batelle Press, 2003.
- P4. Liu B, Sträuber H, Saraiva JP, Harms H, Godinho Silva S, Kleinsteuber S, da Rocha UN. Machine Learning-assisted Identification of Bioindicators Predicts Medium-chain Carboxylate Production Performance of an Anaerobic Mixed Culture. DOI: 10.21203/rs.3.rs-78714/v1
- P3. Saraiva JP, Worrich A, Karakoç C, Kallies R, Chatzinotas A, Centler F, da Rocha UN. Mining synergistic microbial interactions: a roadmap on how to integrate multi-omics data. DOI: 10.20944/preprints202102.0244.v1
- P2. Nuccio EE, Nguyen NH, da Rocha UN, Mayali X, Bougoure J, Weber P, Brodie E, Firestone M, Pett-Ridge J. Community RNA-Seq: Multi-kingdom responses to living versus decaying root inputs in soil. DOI: 10.1101/2021.01.12.426429
- P1. Oliveira Monteiro LM, Saraiva JP, Brizola Toscan R, Stadler PF, Silva-Rocha R, da Rocha UN. PredicTF: a tool to predict bacterial transcription factors in complex microbial communities. DOI: 10.1101/2021.01.28.428666
