Proxies of the Eco-exposome
Motivation
The exposome encompasses an individual's exposure to exogenous chemicals, as well as endogenous chemicals that are produced or altered in response to external stressors. We assume that the extended exposome approach can transferred from human health to ecotoxicology by using a closely related definition with an organism-specific perspective (Escher et al., 2017). Hence, an eco‐exposome approach would require assessing the totality of internal exposure of a specific species using a combined chemical and biological assessment in order to identify toxicologically relevant exposures and to infer adverse outcomes. A required combination of chemical and biological effect assessment is complex and will be faciliated by proxies.
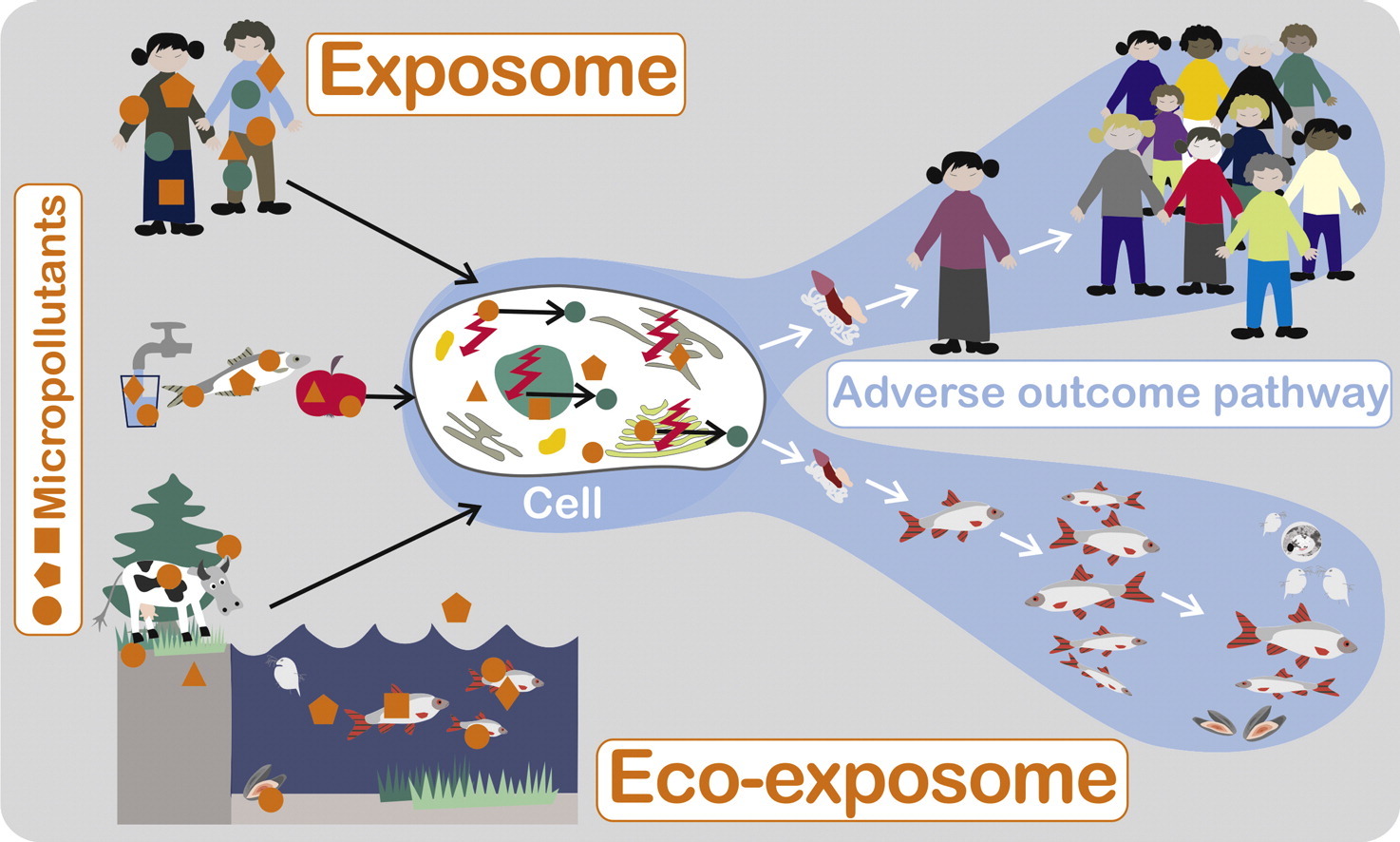
toadverse health outcomes as well as to ecosystem-level effects.
Aims
We aim at a detailed and comprehensive understanding of the fish exposome, in order to infer proxies that could be used for an eco-exposome assessment. We intend to explore whether we can associate complex external to internal signatures and biological responses. Furthermore we intend to identify the best proxies to infer hazard and exposure diagnosis. The project's aim is to contribute to the development of advanced monitoring tools that allow an improved assessment of the environment status.
Structure of the PhD colleg
Coordinator of project:
Stefan Scholz (BIOTOX)
Co-Coordinator of project:
Beate Escher (ZELLTOX)
Team:
Werner Brack (WANA), Wibke Busch (BIOTOX), Paul-Janek Dann (WANA), Jörg Hackermüller (MOLSYB), Annika Jahnke (ZELLTOX), Gianina Jakobs (BIOTOX), Stefan Krämer (MOLSYB), Martin Krauss (WANA), Albrecht Paschke (OEC), Jana Schor (MOLSYB), Theo Wernicke (ZELLTOX)
Four interconnected PhD projects investigate joint samples, which are generated from the same water and fish samples. We aim to contribute to the development of advanced monitoring tools that allow an improved assessment of the environment status. We have access to a set of varying data from internal and external collaboration partners. The four PhD thesis projects aim at
(T1) understanding the relationship of internal and external exposure.
(T2) exploring passive sampling as abiomimetic proxy for the internal exposure
(T3) using untargeted molecular effect analyses in fish embryos for chemical monitoring.
(T4) identifying associations and implementation of suitable data analysis pipelines.
PhD project 4: Link omics data and exposure data
The fourth PhD project is situated in the "Young Investigators Group Bioinformatics and Transcriptomics". Integrating data of exposure and molecular response is at the core of the eco-exposome concept. However, computational methods for this integration and integration with available knowledge are lacking or have not been sufficiently evaluated for this purpose so far. In this PhD project approaches were developed and evaluated to integrate omics data with exposure, using computational methods as like as network inference and association rule mining. The other three PhD projects produce data that can be used to develop and optimize methods and approaches.
We aim to link transcriptional expression effects, internal and external exposure and activity patterns in bio- and chemoassays. This PhD project will generate analysis pipelines, which allow to link exposure and omics data. Furthermore, we aim to develop a deep-learning approach to predict potential adverse outcome pathways out of chemical and biological data sets as well as from knowledge databases.
