Microbial Interaction Ecology
Our group and the UFZ welcome applications, students and collaborators regardless of nationality, religion, gender identification, sexual orientation, age, or disability status. We believe in diverse perspectives and experiences, and thus want to create an environment that helps to find a wide range of potential solutions for scientific questions, but also for society in general.
Research Interests
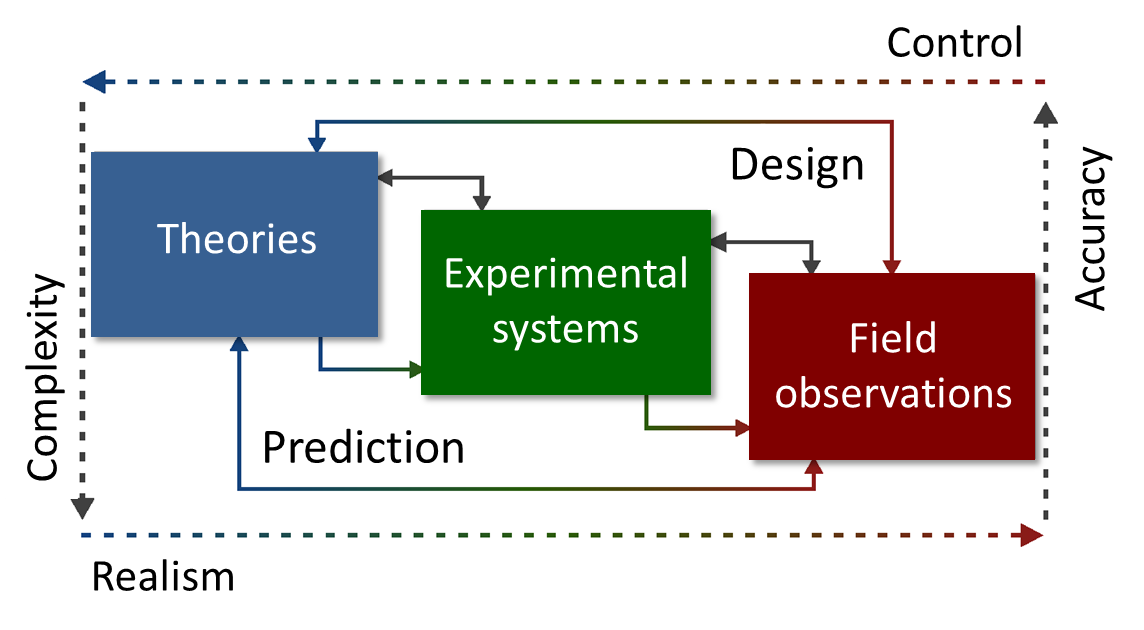
Ecosystems - natural and engineered - are composed of several trophic levels which interact with each other. While bacteria are the key players for many functions, viruses and other micro-predators (protists, bacterial predators) control those communities and thus their functions on multiple levels. Infection by viruses, for instance, has consequences not only for the composition of communities and the genetic landscape but also for biogeochemical turnover und nutrient recycling (e.g., viral shunt).We aim to understand the diversity of viruses and microbes, their interactions and assembly rules, and the consequences for ecosystems by using theory, computational approaches, lab experiments and field surveys.
With this knowledge we hope be able offer solutions for the management of (microbial) ecosystems and the application of viruses/predators in natural, engineered or host-associated systems
Current Projects and Cooperations
- Virus diversity / viromes in soils along land-use gradients (with Michael Schloter, TU Munich, Helmholtz Zentrum Munich; Biodiversity Exploratories SPP 1374)
- Viral shunt, viromes, and virus-host / virus-fungi interactions in subsurface systems and groundwater (with DFG-CRC AquaDiva partners, AquaDiva, UFZ group Bioavailability, Lukas Wick)
- Virus-host interactions: Evolutionary rescue in the context of community complexity, stressors and metacommunities (with Canan Karakoç, Lennon Lab, Indiana Univiersity, and iDiv)
- Microbiomes and viromes in streams (with UFZ Department River Ecology, Markus Weitere)
- Lake and amphibian microbiomes (with Vance Vredenburg , San Francisco State University; ENSAT, Toulouse France; P3)
- Functional traits of phytoplankton along environmental gradients (with iDiv: Susanne Dunker, Stan Harpole, Adam Clark (now University Graz)
- Assessment of pesticide impact on a soil food-web context (with Dimitris Karpouzas, University of Thessaly, Greece, EU-ITN ARISTO)
- The effects of future climatic conditions on protists, predatory microbes and viruses under different land use scenarios (GCEF (with UFZ Department Soil Ecology, iDiv)
- SARS-CoV-2 RNA in wastewater (with TU Dresden, UFZ Groups Attinger, Müller)
Scientists
Felipe Borim Corrêa: Viromics
Dr. Nawras Ghanem: Virus-bacteria interactions, viral shuntDr. Stephanie Jurburg: Microbial Ecology, Community Assembly, Data Stewardship
PhD Students
Peter Hofmann (co-supervision with Susanne Dunker, Stan Harpole): Variability of phytoplankton traits along environmental gradients, microcosm experiments
Marta Eugenia Perez Villanueva (currently in France common PhD student with Cédric Malandain, HYDREKA/Lyon) : Ecotoxicity of pesticides on microbial communities and functions in a food-web context ARISTO Projekt)
Technicians
Anett Heidtmann
Nicole Steinbach
Bachelor and Master students
Guest Scientists
-
Please visit the webpages of the group leader Antonis Chatzinotas or the group members for information on research projects and cooperations.
Publications
To see our publications, please visit our personal web pages.
Find open Postdoc, PhD, Master-/Bachelor-positions and Internships of our group at
Jobs and Education
